Getting Started¶
Estimators in sortedl1 are compatible with the scikit-learn interface. Here is a simple example of fitting a model to some random data.
We start by generating the data.
import numpy as np
from numpy.random import default_rng
from sortedl1 import Slope
# Generate some random data
n = 100
p = 10
seed = 31
rng = default_rng(seed)
x = rng.standard_normal((n, p))
beta = rng.standard_normal(p)
y = x @ beta + rng.standard_normal(n)
Next, we create the estimator by calling Slope() with all the desired parameters.
model = Slope(alpha=0.1)
Now we can fit the model to the data using the fit method, which provides
a fitted model for the given value of alpha above.
model.fit(x, y)
model.coef_
array([ 0.57298856, -1.03709127, 2.55814536, 0.56922823, -1.22227224,
-1.54242725, 0. , -1.30545222, 0. , 0.49147967])
Path Fitting¶
The package also supports fitting the full SLOPE path
via the path method to Slope. In this case, the
value of alpha is ignored and unless path() is called
with a specific sequence of alpha values, a sequence
will automatically be generated to cover solutions from
the point where the first coefficient enters the model
res = model.path(x, y)
Unlike the fit method, calling path() does not modify the model object
and instead returns a named tuple of class PathResults, with the full
set of coefficients and intercepts for each value of alpha.
PathResults also includes concise helpers for quick inspection:
res
res.summary()
{'n_alphas': 77,
'n_features': 10,
'n_targets': 1,
'alpha_min': 0.0010078421223305043,
'alpha_max': 1.1860406557259944,
'coef_shape': (10, 1, 77),
'intercepts_shape': (1, 77),
'lambda_shape': (10,),
'nnz_first_alpha': 0,
'nnz_last_alpha': 10}
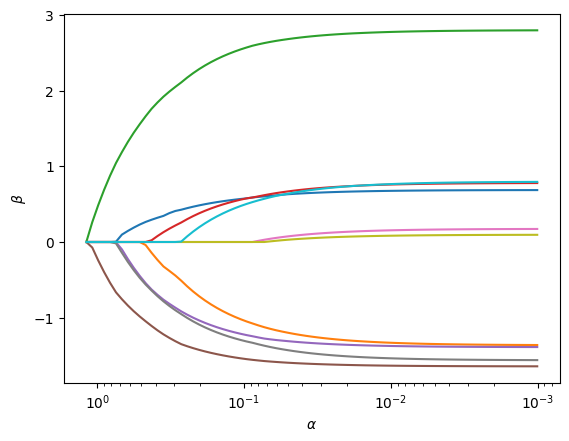
It also comes with a plot() method to visualize the path of coefficients:
fig, ax = res.plot()
Cross-Validation¶
It is also easy to cross-validate in the sortedl1 package. Since the estimator is scikit-learn compatible, we could use the functionality from scikit-learn directly, but sortedl1 also includes native cross-validation routines that are optimized for the SLOPE package.
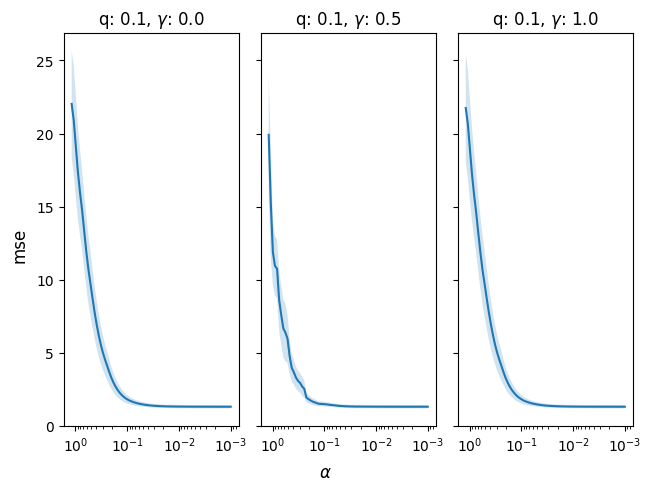
In the following example, we cross-validate
across different levels of the gamma parameter,
which fits the relaxed SLOPE model (a linear combination
of SLOPE and ordinary least squares fit to the
cluster structure from SLOPE).
cv_res = model.cv(x, y, q=[0.1], gamma=[0.0,0.5, 1.0])
fig, ax = cv_res.plot()
cv_res
cv_res.summary()
{'metric': 'mse',
'n_param_sets': 3,
'best_ind': 1,
'best_alpha_ind': 76,
'best_score': 1.0,
'best_alpha': 0.0010078421223305043,
'n_alphas_per_param': [77, 77, 77],
'param_keys': ['gamma', 'q']}
If you want both cross-validation results and a best model fitted on the full
data used in cross-validation, use refit=True:
cv_res, best_model = model.cv(x, y, q=[0.1], gamma=[0.0, 0.5, 1.0], refit=True)
best_model.coef_
array([ 0.67405566, -1.32637578, 2.77084901, 0.75603046, -1.37102155,
-1.63196265, 0.15177179, -1.53242048, 0.08127653, 0.76057772])
In this low-dimensional example, we see that there is, unsurprisingly, little benefit to regularization.
Using scikit-learn model selection¶
The Slope estimator can also be used directly with scikit-learn
model-selection tools such as GridSearchCV.
from sklearn.model_selection import GridSearchCV
param_grid = {"alpha": [0.01, 0.1, 1.0], "q": [0.05, 0.1, 0.2]}
search = GridSearchCV(Slope(), param_grid=param_grid, cv=5)
search.fit(x, y)
search.best_params_
---------------------------------------------------------------------------
TypeError Traceback (most recent call last)
Cell In[9], line 5
1 from sklearn.model_selection import GridSearchCV
2
3 param_grid = {"alpha": [0.01, 0.1, 1.0], "q": [0.05, 0.1, 0.2]}
4 search = GridSearchCV(Slope(), param_grid=param_grid, cv=5)
----> 5 search.fit(x, y)
6
7 search.best_params_
File ~/.local/lib/python3.12/site-packages/sklearn/base.py:1336, in _fit_context.<locals>.decorator.<locals>.wrapper(estimator, *args, **kwargs)
1329 estimator._validate_params()
1331 with config_context(
1332 skip_parameter_validation=(
1333 prefer_skip_nested_validation or global_skip_validation
1334 )
1335 ):
-> 1336 return fit_method(estimator, *args, **kwargs)
File ~/.local/lib/python3.12/site-packages/sklearn/model_selection/_search.py:955, in BaseSearchCV.fit(self, X, y, **params)
921 """Run fit with all sets of parameters.
922
923 Parameters
(...) 952 Instance of fitted estimator.
953 """
954 estimator = self.estimator
--> 955 scorers, refit_metric = self._get_scorers()
957 X, y = indexable(X, y)
958 params = _check_method_params(X, params=params)
File ~/.local/lib/python3.12/site-packages/sklearn/model_selection/_search.py:852, in BaseSearchCV._get_scorers(self)
850 scorers = self.scoring
851 elif self.scoring is None or isinstance(self.scoring, str):
--> 852 scorers = check_scoring(self.estimator, self.scoring)
853 else:
854 scorers = _check_multimetric_scoring(self.estimator, self.scoring)
File ~/.local/lib/python3.12/site-packages/sklearn/utils/_param_validation.py:218, in validate_params.<locals>.decorator.<locals>.wrapper(*args, **kwargs)
212 try:
213 with config_context(
214 skip_parameter_validation=(
215 prefer_skip_nested_validation or global_skip_validation
216 )
217 ):
--> 218 return func(*args, **kwargs)
219 except InvalidParameterError as e:
220 # When the function is just a wrapper around an estimator, we allow
221 # the function to delegate validation to the estimator, but we replace
222 # the name of the estimator by the name of the function in the error
223 # message to avoid confusion.
224 msg = re.sub(
225 r"parameter of \w+ must be",
226 f"parameter of {func.__qualname__} must be",
227 str(e),
228 )
File ~/.local/lib/python3.12/site-packages/sklearn/metrics/_scorer.py:996, in check_scoring(estimator, scoring, allow_none, raise_exc)
994 return None
995 else:
--> 996 raise TypeError(
997 "If no scoring is specified, the estimator passed should "
998 "have a 'score' method. The estimator %r does not." % estimator
999 )
TypeError: If no scoring is specified, the estimator passed should have a 'score' method. The estimator Slope() does not.