An introduction to qualpalr
Johan Larsson
2026-01-30
Source:vignettes/introduction.Rmd
introduction.RmdOverview
qualpalr generates qualitative color palettes optimized
for maximally distinct colors. Given n (the number of
colors to generate), along with a subset in the hsl color space1 (a
cylindrical representation of the RGB color space) qualpalr
attempts to find the n colors in the provided color
subspace that maximize the smallest pairwise color difference.
This is done by computing the pairwise color differences between all the
input colors, and then selecting the n colors that maximize
the minimum pairwise color difference.
Examples
qualpalr main workhorse is qualpal(), which
takes as its input n (the number of colors to generate) and
colorspace, which can be either
- a list of numeric vectors
h(hue from -360 to 360),s(saturation from 0 to 1), andl(lightness from 0 to 1), all of length 2, specifying a min and max, - a list of numeric vectors
h(hue from -360 to 360),c(chroma from 0 to 100), andl(lightness from 0 to 100), all of length 2, specifying a min and max, or - a character vector specifying a predefined color palette.
library(qualpalr)
pal <- qualpal(5, list(h = c(-200, 120), s = c(0.3, 0.8), l = c(0.4, 0.9)))
# Adapt the color space to deuteranopia of severity 0.7
pal <- qualpal(5, cvd = c(deutan = 0.7))The resulting object, pal, is a list with several color
tables and a distance matrix based based on the color difference metric
used, by default CIEDE2000 (metric = ciede2000).
pal## ----------------------------------------
## Colors in the HSL color space
##
## Hue Saturation Lightness
## #ca6c74 355 0.47 0.61
## #6e6cca 241 0.47 0.61
## #c6a5db 277 0.43 0.75
## #c7eadc 155 0.46 0.85
## #c9cb70 62 0.47 0.62
##
## ----------------------------------------
## DIN99d color difference distance matrix
##
## #ca6c74 #6e6cca #c6a5db #c7eadc
## #6e6cca 22
## #c6a5db 17 14
## #c7eadc 25 25 20
## #c9cb70 22 29 24 16Methods for pairs and plot have been
written for qualpal objects to help visualize the
results.
# Multidimensional scaling plot
plot(pal)
# Pairs plot in the DIN99d color space
pairs(pal, colorspace = "DIN99d")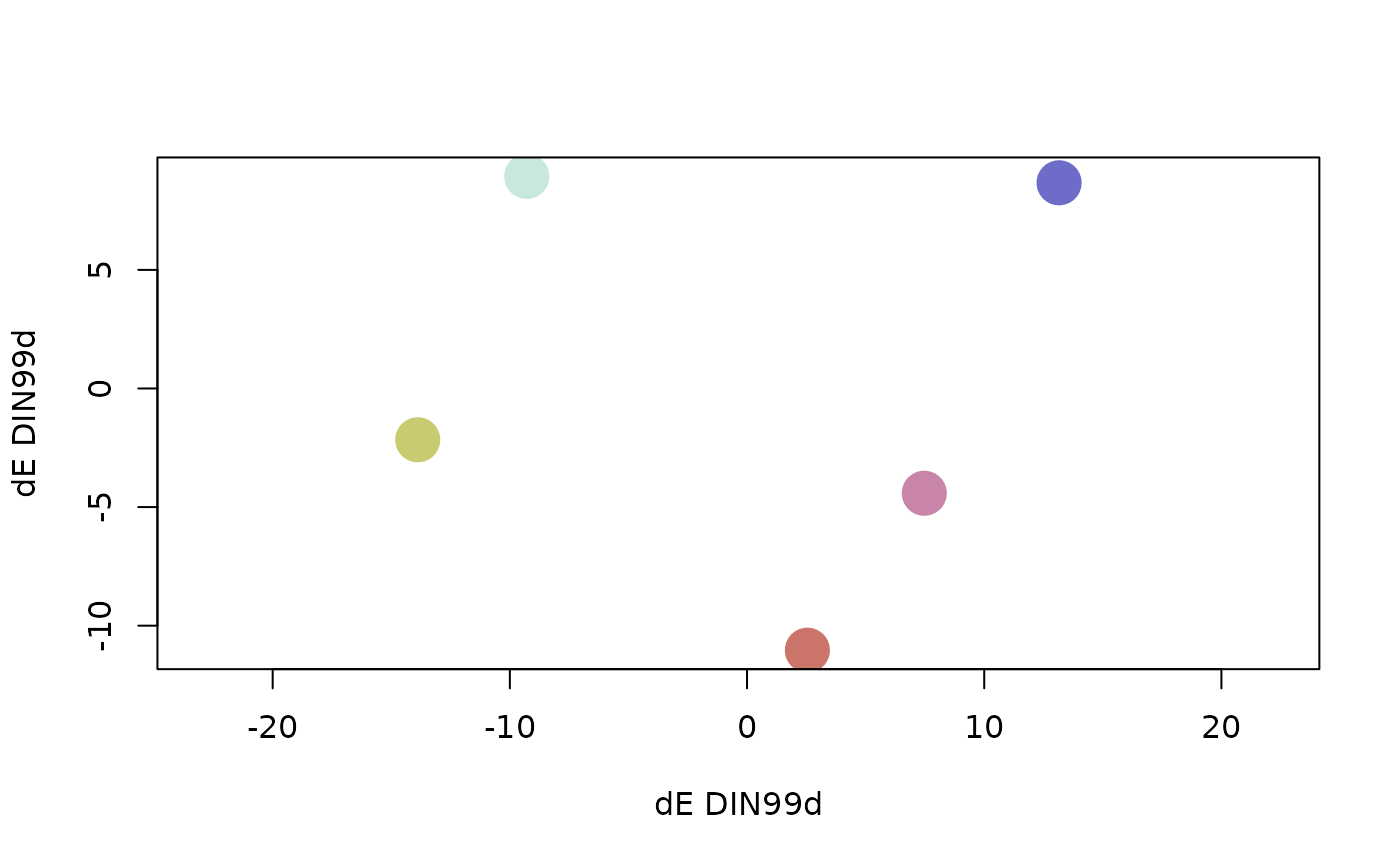
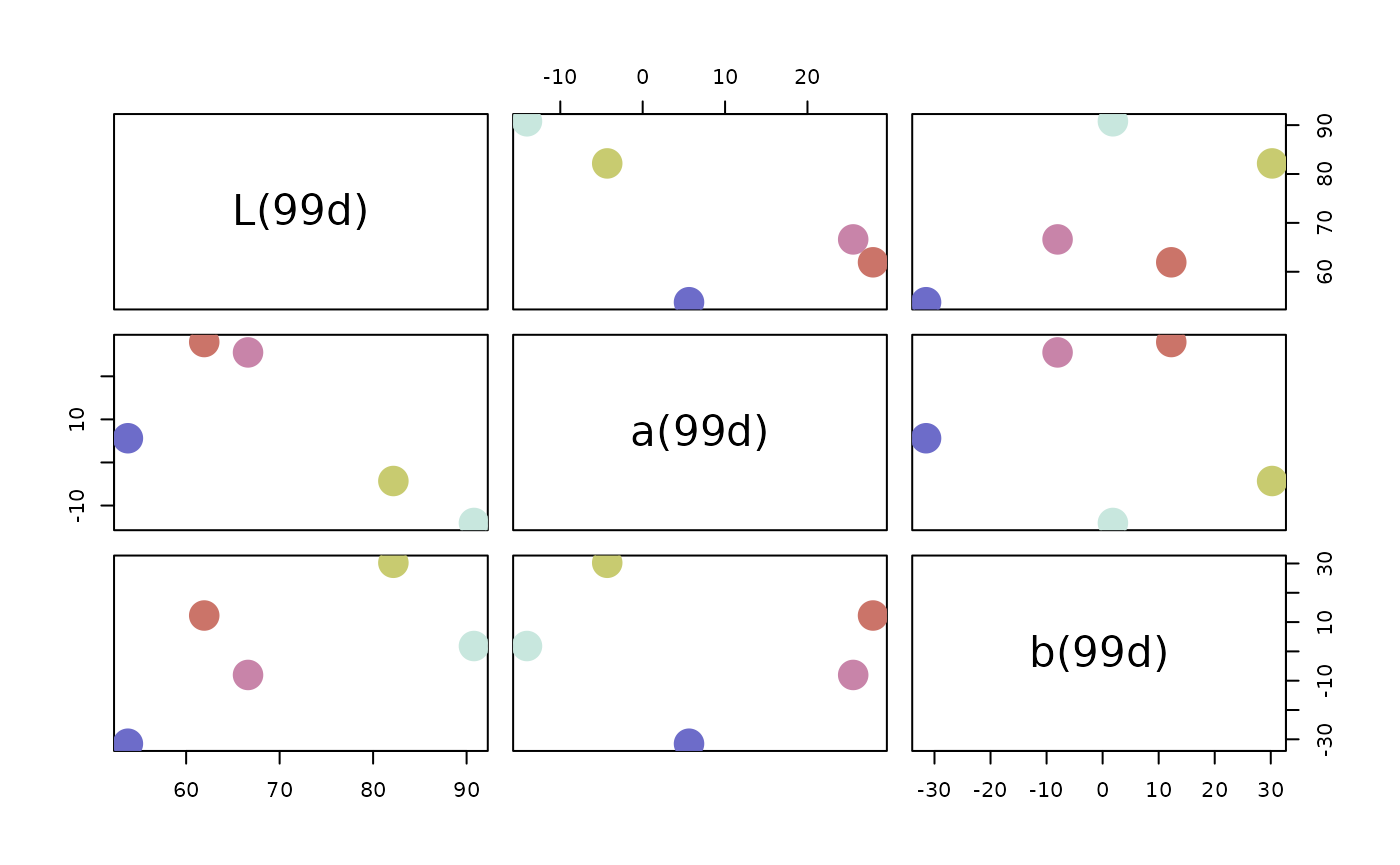

The colors are most easily used in R by accessing
pal$hex
Details
qualpal begins by generating a point cloud out of the
HSL color subspace provided by the user, using a quasi-random Halton
sequence. Here is the color subspace in HSL with settings
h = c(-200, 120), s = c(0.3, 0.8), l = c(0.4, 0.9).
The program then proceeds by projecting these colors into the sRGB space.
It then continues projecting the colors into the XYZ space. After this, behavior depends on the metric used. By default, qualpal uses the CIEDE2000 color difference formula (Sharma, Wu, and Dalal 2005), which is the current state of the art in color difference metrics and standard as defined by the International Commission on Illumination (CIE). For illustrative purposes, however, we will show the procedure when the DIN99d color space (Cui et al. 2002) is used instead, which is a perceptually uniform color space that uses the Euclidean distance as a color difference metric. This makes for a computationally simpler and faster algorithm, but it is not as accurate as CIEDE2000.
When using the DIN99d color space, we also apply a power transformation (Huang et al. 2015) to fine tune these differences.
To select the n colors that the user wanted, we proceed
greedily: first, we find the two most distant points, then we find the
third point that maximizes the minimum distance to the previously
selected points. This is repeated until n points are
selected. These points are then returned to the user; below is an
example using n = 5.
Thanks
Bruce Lindbloom’s webpage has been instrumental in making qualpalr. Thanks also to i want hue, which inspired me to make qualpalr.