
An introduction to SLOPE
Johan Larsson
2026-03-30
Source:vignettes/introduction.Rmd
introduction.RmdBackground
The functions in this package solves problems of the type where the second part of the objective is the sorted L1-norm where , and represents an rank of the magnitudes of in descending order. controls the shape of the penalty sequence, which needs to be non-increasing, and controls the scale of that sequence.
Solving this problem is called SLOPE (Sorted L-One Penalized Estimation) (Bogdan et al. 2015).
In this problem, is a smooth and convex objective, which for this package so far includes four models from the family of generalized linear models:
- Gaussian regression,
- binomial regression,
- multinomial regression, and
- Poisson regression.
SLOPE is an extension of the lasso (Tibshirani 1996) and has the ability to lead to sparse solutions given a sufficiently strong regularization. It is also easy to see that SLOPE reduces to the lasso if all elements of the vector are equal.
The lasso, however, has difficulties with correlated predictors (Jia and Yu 2010) but this is not the case with SLOPE, which handles this issue by clustering predictors to the same magnitude. This effect is related to the consecutive differences of the vector: the larger the steps, the more clustering behavior SLOPE exhibits.
An example
In the following example, we will use the heart data set, for which
the response is a cardiac event. (The package contains several data sets
to exemplify modeling. Please see the examples in SLOPE().)
The main function of the package is SLOPE(), which, more or
less, serves as an interface for code written in C++. There are many
arguments in the function and most of them relate to either the
construction of the regularization path or the penalty
()
sequence used. Here we will use the option lambda = "bh",
which used the BH method detailed in Bogdan et
al. (2015) to select the sequence. (Note that it is also possible
to manually insert a sequence.)
The default print method gives a summary of the regularization path but it is usually more informative to study a plot of the path.
plot(fit)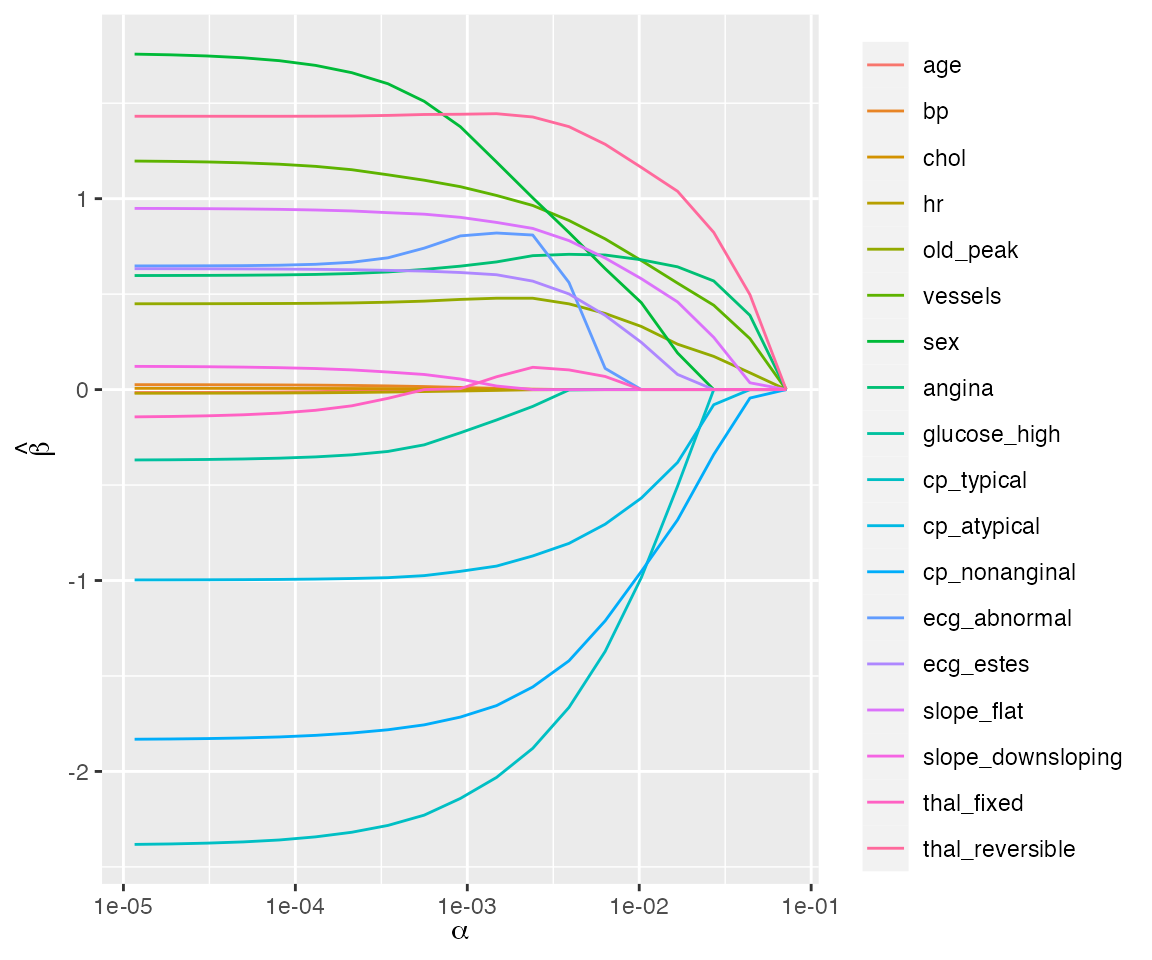
Regularization path for a binomial regression model fit to the heart data set.
Cross-validation
To determine the strength of regularization, it is almost always
necessary to tune the
sequence using resampling. This package features the function
trainSLOPE() to do this. We will give an example of
trainSLOPE() here.
set.seed(924)
x <- bodyfat$x
y <- bodyfat$y
tune <- trainSLOPE(
x,
y,
q = c(0.1, 0.2),
number = 5,
repeats = 2
)As before, the plot method offers the best summary.
plot(tune, measure = "mae") # plot mean absolute error
#> Warning: `measure` is deprecated, and will be removed in a future version. the
#> measure will instead be taken from the `TrainedSLOPE` objectPrinting the resulting object will display the optimum values
tune
#>
#> Call:
#> trainSLOPE(x = x, y = y, q = c(0.1, 0.2), number = 5, repeats = 2)
#>
#> Optimum values:
#> q alpha measure mean se lo hi
#> 1 0.2 0.001638533 mae 3.613623 0.1319657 3.315096 3.912151
#> 2 0.2 0.002166044 mse 19.535805 1.1247987 16.991333 22.080276False discovery rate
Under assumptions of orthonormality, SLOPE has been shown to control false discovery rate (FDR) of non-zero coefficients (feature weights) in the model (Bogdan et al. 2015). It is in many ways analogous to the Benjamini–Hochberg procedure for multiple comparisons.
Let’s set up a simple experiment to see how SLOPE controls the FDR. We randomly generate data sets with various proportions of true signals. Under this Gaussian design with independently and identically distributed columns in , SLOPE should asymptotically control FDR at the level given by the shape parameter , which we set to 0.1 in this example.
# proportion of real signals
q <- seq(0.05, 0.5, length.out = 20)
fdr <- double(length(q))
set.seed(1)
for (i in seq_along(q)) {
n <- 1000
p <- n / 2
alpha <- 1
problem <- SLOPE:::randomProblem(n, p, q = q[i], alpha = alpha)
x <- problem$x
y <- problem$y
signals <- problem$nonzero
fit <- SLOPE(x,
y,
lambda = "gaussian",
q = 0.1,
alpha = alpha / sqrt(n)
)
selected_slope <- which(as.matrix(fit$nonzeros[[1]]))
v <- length(setdiff(selected_slope, signals))
r <- length(selected_slope)
fdr[i] <- v / max(r, 1)
}
plot(
q,
fdr,
type = "o",
xlab = "q",
ylab = "FDR",
pch = 16,
las = 1
)
abline(h = 0.1, lty = 3)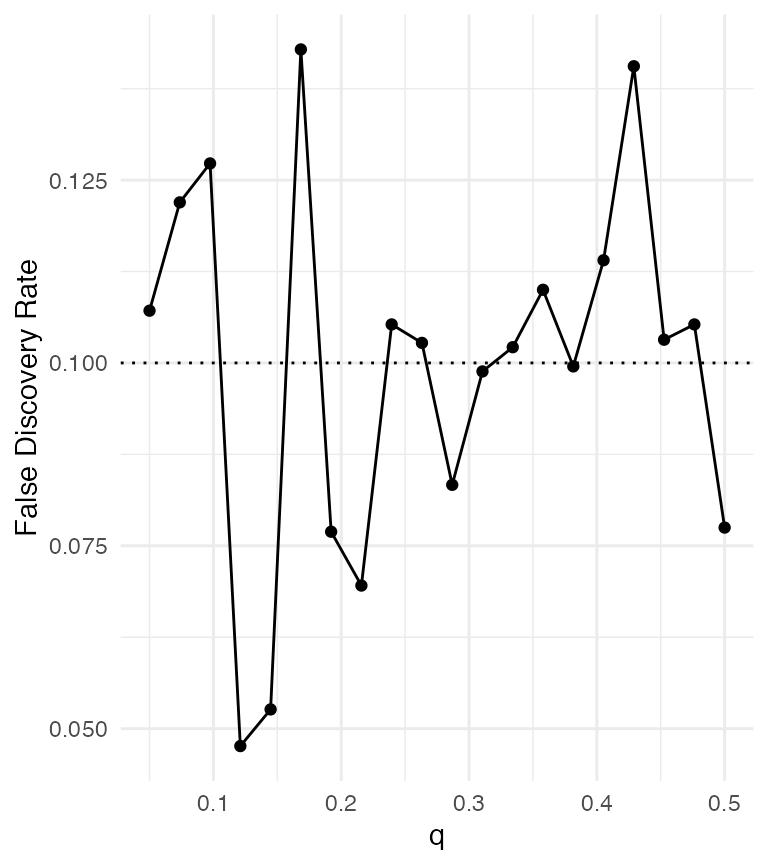
Control of false discovery rate using SLOPE.
SLOPE seems to control FDR at roughly the specified level.